Note
Go to the end to download the full example code.
Define a space-varying cell model#
This example shows how to define a space-varying cell model for the ventricles
and septum of a full-heart model. The cell model parameters (gks and gto)
are linearly interpolated across a 2-D grid defined by the endo-epicardial and
apico-basal directions.
Perform the required imports#
Import the required modules and set relevant paths.
from pathlib import Path
import numpy as np
import pyvista as pv
from ansys.health.heart.examples import get_preprocessed_fullheart
import ansys.health.heart.models as models
import ansys.health.heart.settings.material.cell_models as cell_models
Load the full-heart model#
Set the working directory, download the preprocessed model and load it.
workdir = Path.home() / "pyansys-heart" / "downloads" / "Rodero2021" / "01" / "FullHeart"
path_to_model, path_to_partinfo, _ = get_preprocessed_fullheart(resolution="2.0mm")
model: models.FullHeart = models.FullHeart.load_model(
path_to_model, path_to_partinfo, working_directory=workdir
)
Normalize the coordinate fields#
Extract and normalize the transmural (endo-epi) and apico-basal fields to [0, 1].
endo_epi = model.mesh["transmural"].copy()
apex_base = model.mesh["apico-basal"].copy()
endo_epi = (endo_epi - np.nanmin(endo_epi)) / (np.nanmax(endo_epi) - np.nanmin(endo_epi))
apex_base = (apex_base - np.nanmin(apex_base)) / (np.nanmax(apex_base) - np.nanmin(apex_base))
Define helper functions#
define_node_group bins nodes by two normalized coordinate fields.
def define_node_group(selected_nodes, coord1, coord2, bins1=[0, 1], bins2=[0, 1]):
"""Define node groups based on two coordinates and their respective bins.
Parameters
----------
selected_nodes : array-like
Indices of the selected nodes.
coord1 : array-like
First coordinate values for the nodes.
coord2 : array-like
Second coordinate values for the nodes.
bins1 : list, default: [0, 1]
Bin edges for the first coordinate.
bins2 : list, default: [0, 1]
Bin edges for the second coordinate.
Returns
-------
list of list of np.ndarray
Node groups organized by the bins of coord1 and coord2.
"""
masked_coord1 = coord1.copy()
masked_coord2 = coord2.copy()
# set nan for the rest so that they will not be included in the group definition
masked_coord1[~np.isin(np.arange(len(masked_coord1)), selected_nodes)] = np.nan
masked_coord2[~np.isin(np.arange(len(masked_coord2)), selected_nodes)] = np.nan
node_groups = [[None] * (len(bins2) - 1) for _ in range(len(bins1) - 1)]
for i in range(len(bins1) - 1):
for j in range(len(bins2) - 1):
mask = (
(masked_coord1 >= bins1[i])
& (masked_coord1 <= bins1[i + 1])
& (masked_coord2 >= bins2[j])
& (masked_coord2 <= bins2[j + 1])
)
node_groups[i][j] = np.where(mask)[0]
return node_groups
Define the binning and parameter ranges#
Set the bin edges for the endo-epi and apex-base directions, and the
extremal values of gks and gto used for linear interpolation.
endo_epi_bins = [0, 0.17, 0.41, 1] # 0-0.17: endo, 0.17-0.41: mid, 0.41-1: epi
apex_base_bins = [0, 0.5, 0.65, 0.8, 1] # apex to base binned into 4 groups
# Set bounds for gks and gto based on the range of values in Tentusscher cell models,
# which will be used for 2D linear interpolation.
gks_min = cell_models.TentusscherEndo().gks # extreme value at endo apex
gks_max = cell_models.TentusscherEpi().gks # extreme value at epi base
gto_min = cell_models.TentusscherEndo().gto # extreme value at endo apex
gto_max = cell_models.TentusscherEpi().gto # extreme value at epi base
Assign cell models to the ventricles#
Group left- and right-ventricle nodes by endo-epi and apex-base bins, then
interpolate gks and gto linearly across the 2-D grid.
lv_nodes = model.left_ventricle.get_node_ids(model.mesh)
rv_nodes = model.right_ventricle.get_node_ids(model.mesh)
ventricle_nodes = np.unique(np.concatenate((lv_nodes, rv_nodes)))
ventricle_node_groups = define_node_group(
ventricle_nodes, endo_epi, apex_base, endo_epi_bins, apex_base_bins
)
n_row, n_col = len(endo_epi_bins) - 1, len(apex_base_bins) - 1
factors = np.arange(n_row * n_col).reshape(n_row, n_col) / (n_row * n_col - 1)
gks_matrix = gks_min + factors * (gks_max - gks_min)
gto_matrix = gto_min + factors * (gto_max - gto_min)
ventricle_cell_models = [
[cell_models.Tentusscher(gks=gks_matrix[i, j], gto=gto_matrix[i, j]) for j in range(n_col)]
for i in range(n_row)
]
ventricle_groups_flat = [x for sublist in ventricle_node_groups for x in sublist]
ventricle_models_flat = [x for sublist in ventricle_cell_models for x in sublist]
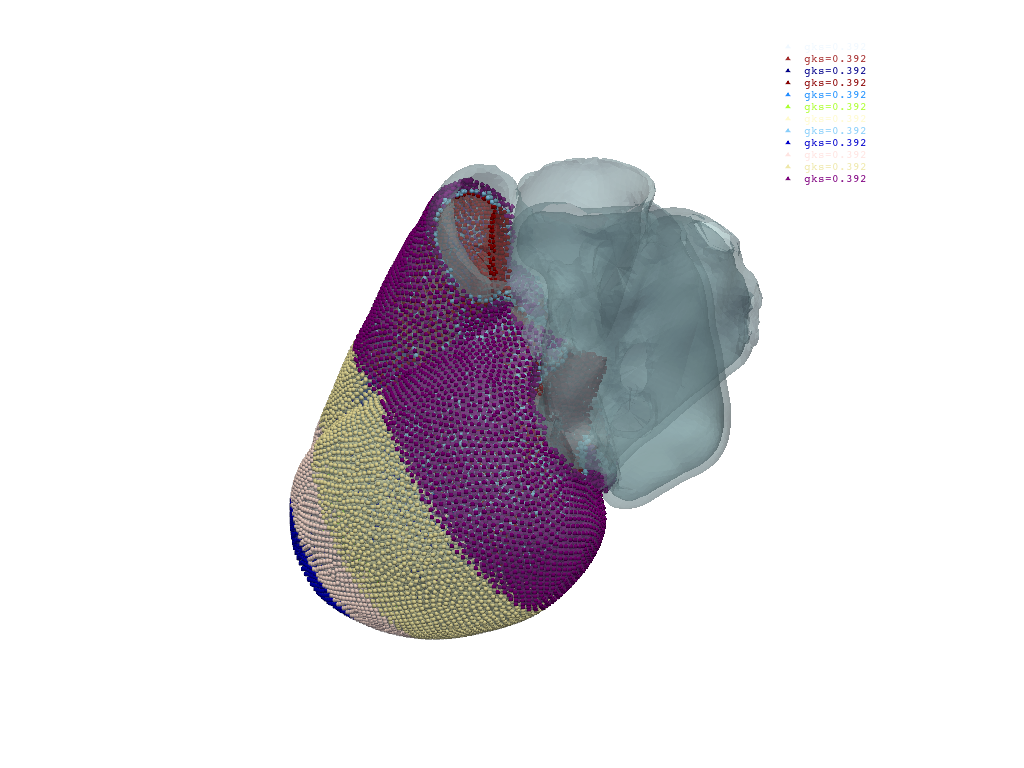
Plot ventricular node groups#
Each group is rendered with a distinct color and labeled with its gks value.
colors = list(pv.colors.hexcolors.keys())[::10]
p = pv.Plotter()
model.mesh.set_active_scalars(None)
p.add_mesh(model.mesh, opacity=0.2)
for i in range(len(ventricle_groups_flat)):
point_cloud = pv.PolyData(model.mesh.points[ventricle_groups_flat[i]])
color = colors[i % len(colors)]
p.add_mesh(
point_cloud,
color=color,
label=f"gks={ventricle_models_flat[i].gks:.3f}",
point_size=5,
render_points_as_spheres=True,
)
p.add_legend()
p.show()

Assign cell models to the septum#
For the septum, only the apex-base direction is used for variation. Parameters will be interpolated from endo values.
septum_nodes = model.septum.get_node_ids(model.mesh)
septum_node_groups = define_node_group(septum_nodes, endo_epi, apex_base, bins2=apex_base_bins)
n_row, n_col = 1, len(apex_base_bins) - 1
factors = np.arange(n_row * n_col).reshape(n_row, n_col) / (n_row * n_col - 1)
factors = np.array([[0]]) if n_row * n_col == 1 else factors # handle the case with only one group
gks_matrix = gks_min + factors * (gks_max - gks_min)
gto_matrix = gto_min + factors * (gto_max - gto_min)
septum_cell_models = [
[cell_models.Tentusscher(gks=gks_matrix[i, j], gto=gto_matrix[i, j]) for j in range(n_col)]
for i in range(n_row)
]
septum_groups_flat = [x for sublist in septum_node_groups for x in sublist]
septum_models_flat = [x for sublist in septum_cell_models for x in sublist]
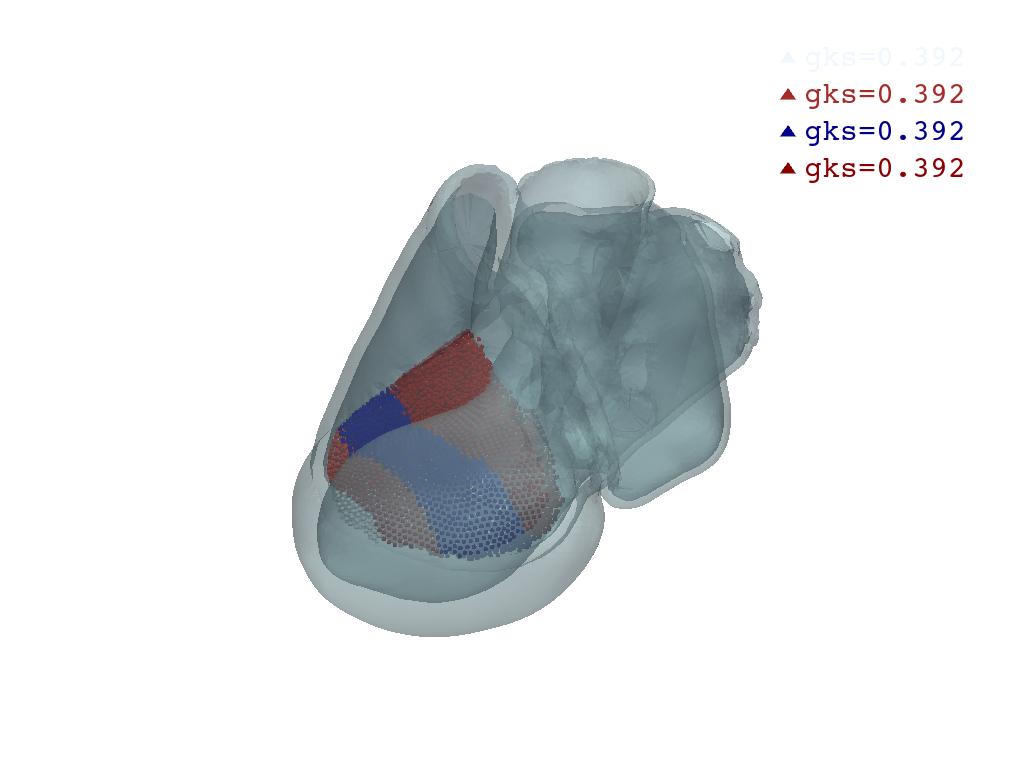
Plot septum node groups#
p = pv.Plotter()
model.mesh.set_active_scalars(None)
p.add_mesh(model.mesh, opacity=0.2)
for i in range(len(septum_groups_flat)):
point_cloud = pv.PolyData(model.mesh.points[septum_groups_flat[i]])
color = colors[i % len(colors)]
p.add_mesh(
point_cloud,
color=color,
label=f"gks={septum_models_flat[i].gks:.3f}",
point_size=5,
render_points_as_spheres=True,
)
p.add_legend()
p.show()
Merge and assign all cell models#
Combine the ventricle and septum groups and models into flat lists ready for assignment to the heart model.
all_groups_flat = ventricle_groups_flat + septum_groups_flat
all_models_flat = ventricle_models_flat + septum_models_flat
# The cell model will be defined based on the nodeset
# This overwrites any existing cell model assignment based on part.
model._nodeset_cellmodel = (all_groups_flat, all_models_flat)
Total running time of the script: (0 minutes 13.225 seconds)

