Note
Go to the end to download the full example code.
Create a full-heart model#
This example shows how to process a case from Rodero et al. (2021) into a simulation-ready heart model.
Perform the required imports#
Import the required modules and set relevant paths, including that of the working directory and generated model.
import json
import os
from pathlib import Path
import ansys.health.heart.models as models
from ansys.health.heart.pre.database_utils import get_compatible_input
from ansys.health.heart.utils.download import download_case_from_zenodo, unpack_case
# specify a download directory
download_folder = Path.home() / "pyansys-heart" / "downloads"
# Download a compatible case from the Zenodo database.
tar_file = download_case_from_zenodo("Rodero2021", 1, download_folder, overwrite=False)
# Unpack the case to get the input CASE or VTK file.
case_file = unpack_case(tar_file)
# Specify the working directory. This code uses the directory of the CASE file.
workdir = os.path.join(os.path.dirname(case_file), "FullHeart")
if not os.path.isdir(workdir):
os.makedirs(workdir)
# Specify paths to the model, input, and part definitions.
path_to_model = os.path.join(workdir, "heart_model.vtu")
path_to_input = os.path.join(workdir, "input_model.vtp")
path_to_part_definitions = os.path.join(workdir, "part_definitions.json")
Note
You can also manually download the CASE or VTK files from the Strocchi 2020 and Rodero 2021 databases. For more information, see:
Alternatively, you can simply click one of the buttons at the bottom of this page to download a CASE file for the Rodero 2021 database in an IPYNB, PY, or ZIP format.
Convert the VTK file to a compatible input format#
input_geom, part_definitions = get_compatible_input(
case_file, model_type="FullHeart", database="Rodero2021"
)
# Note that the input model and part definitions can be saved for later use.
# Save input geometry and part definitions.
input_geom.save(path_to_input)
with open(path_to_part_definitions, "w") as f:
json.dump(part_definitions, f, indent=True)
Create a heart model#
Create a full-heart model.
model = models.FullHeart(working_directory=workdir)
# Load input model generated in an earlier step.
model.load_input(input_geom, part_definitions, "surface-id")
# Mesh the volume of all structural parts.
model.mesh_volume(use_wrapper=True, global_mesh_size=2.0, _global_wrap_size=2.0)
# Update the model and extract the required anatomical features.
model.update()
# Optionally save the simulation mesh as a VTK object for "offline" inspection.
model.mesh.save(os.path.join(model.workdir, "simulation-mesh.vtu"))
model.save_model(os.path.join(model.workdir, "heart_model.vtu"))
# Print some information about the processed model.
print(model)
# Print part names.
print(model.part_names)
C:\Users\ansys\actions-runner\_work\pyansys-heart\pyansys-heart\.tox\doc-html\Lib\site-packages\ansys\health\heart\utils\vtk_utils.py:149: PyVistaDeprecationWarning: This filter is deprecated. Use `select_interior_points` instead.
centroids = centroids.select_enclosed_points(surface, tolerance=tolerance, check_surface=True)
C:\Users\ansys\actions-runner\_work\pyansys-heart\pyansys-heart\.tox\doc-html\Lib\site-packages\ansys\health\heart\utils\vtk_utils.py:149: PyVistaDeprecationWarning: This filter is deprecated. Use `select_interior_points` instead.
centroids = centroids.select_enclosed_points(surface, tolerance=tolerance, check_surface=True)
C:\Users\ansys\actions-runner\_work\pyansys-heart\pyansys-heart\.tox\doc-html\Lib\site-packages\ansys\health\heart\utils\vtk_utils.py:149: PyVistaDeprecationWarning: This filter is deprecated. Use `select_interior_points` instead.
centroids = centroids.select_enclosed_points(surface, tolerance=tolerance, check_surface=True)
C:\Users\ansys\actions-runner\_work\pyansys-heart\pyansys-heart\.tox\doc-html\Lib\site-packages\ansys\health\heart\utils\vtk_utils.py:149: PyVistaDeprecationWarning: This filter is deprecated. Use `select_interior_points` instead.
centroids = centroids.select_enclosed_points(surface, tolerance=tolerance, check_surface=True)
C:\Users\ansys\actions-runner\_work\pyansys-heart\pyansys-heart\.tox\doc-html\Lib\site-packages\ansys\health\heart\utils\vtk_utils.py:149: PyVistaDeprecationWarning: This filter is deprecated. Use `select_interior_points` instead.
centroids = centroids.select_enclosed_points(surface, tolerance=tolerance, check_surface=True)
C:\Users\ansys\actions-runner\_work\pyansys-heart\pyansys-heart\.tox\doc-html\Lib\site-packages\ansys\health\heart\utils\vtk_utils.py:149: PyVistaDeprecationWarning: This filter is deprecated. Use `select_interior_points` instead.
centroids = centroids.select_enclosed_points(surface, tolerance=tolerance, check_surface=True)
C:\Users\ansys\actions-runner\_work\pyansys-heart\pyansys-heart\.tox\doc-html\Lib\site-packages\ansys\health\heart\utils\vtk_utils.py:149: PyVistaDeprecationWarning: This filter is deprecated. Use `select_interior_points` instead.
centroids = centroids.select_enclosed_points(surface, tolerance=tolerance, check_surface=True)
GENERAL:
total_num_tets: 331365
total_num_nodes: 72171
PARTS:
Left ventricle:
num_tets: 140461
SURFACES:
Left ventricle endocardium:
num_faces: 7421
Left ventricle epicardium:
num_faces: 8126
Left ventricle septum:
num_faces: 0
CAPS:
mitral-valve:
num_nodes: 67
aortic-valve:
num_nodes: 44
Right ventricle:
num_tets: 67570
SURFACES:
Right ventricle endocardium:
num_faces: 8875
Right ventricle epicardium:
num_faces: 9339
Right ventricle septum:
num_faces: 2758
CAPS:
tricuspid-valve:
num_nodes: 93
pulmonary-valve:
num_nodes: 58
Septum:
num_tets: 42589
SURFACES: {}
CAPS: {}
Left atrium:
num_tets: 31533
SURFACES:
Left atrium endocardium:
num_faces: 5740
Left atrium epicardium:
num_faces: 4373
CAPS:
mitral-valve-atrium:
num_nodes: 45
right-superior-pulmonary-vein:
num_nodes: 37
left-superior-pulmonary-vein:
num_nodes: 35
right-inferior-pulmonary-vein:
num_nodes: 31
left-atrium-appendage:
num_nodes: 28
left-inferior-pulmonary-vein:
num_nodes: 26
Right atrium:
num_tets: 32089
SURFACES:
Right atrium endocardium:
num_faces: 7168
Right atrium epicardium:
num_faces: 5574
CAPS:
tricuspid-valve-atrium:
num_nodes: 73
inferior-vena-cava:
num_nodes: 35
superior-vena-cava:
num_nodes: 34
Aorta:
num_tets: 12731
SURFACES:
Aorta wall:
num_faces: 4290
CAPS: {}
Pulmonary artery:
num_tets: 4392
SURFACES:
Pulmonary artery wall:
num_faces: 1348
CAPS: {}
CAVITIES:
Left ventricle cavity:
volume: 120446.96049660796
Right ventricle cavity:
volume: 185648.45053118863
Left atrium cavity:
volume: 68001.90309648326
Right atrium cavity:
volume: 110948.1886515331
['Left ventricle', 'Right ventricle', 'Septum', 'Left atrium', 'Right atrium', 'Aorta', 'Pulmonary artery']
Visualize results#
Visualize and inspect the components of the model by accessing various properties or attributes and invoking methods.
print(f"Volume of LV cavity: {model.left_ventricle.cavity.volume} mm^3")
print(f"Volume of LV cavity: {model.left_atrium.cavity.volume} mm^3")
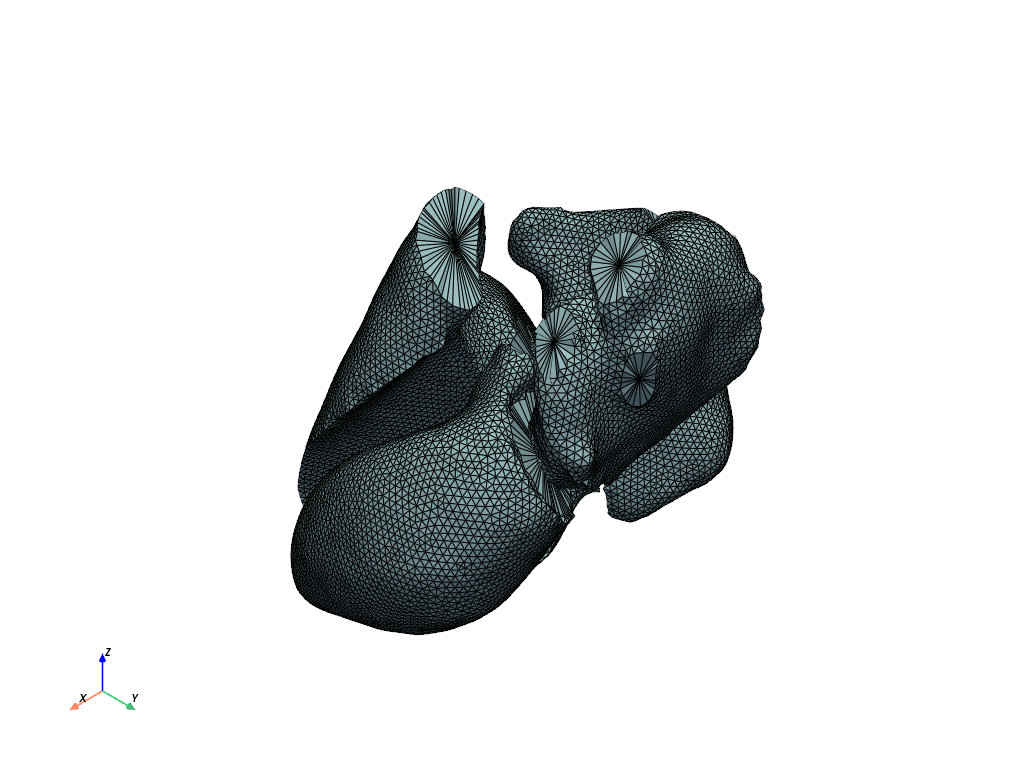
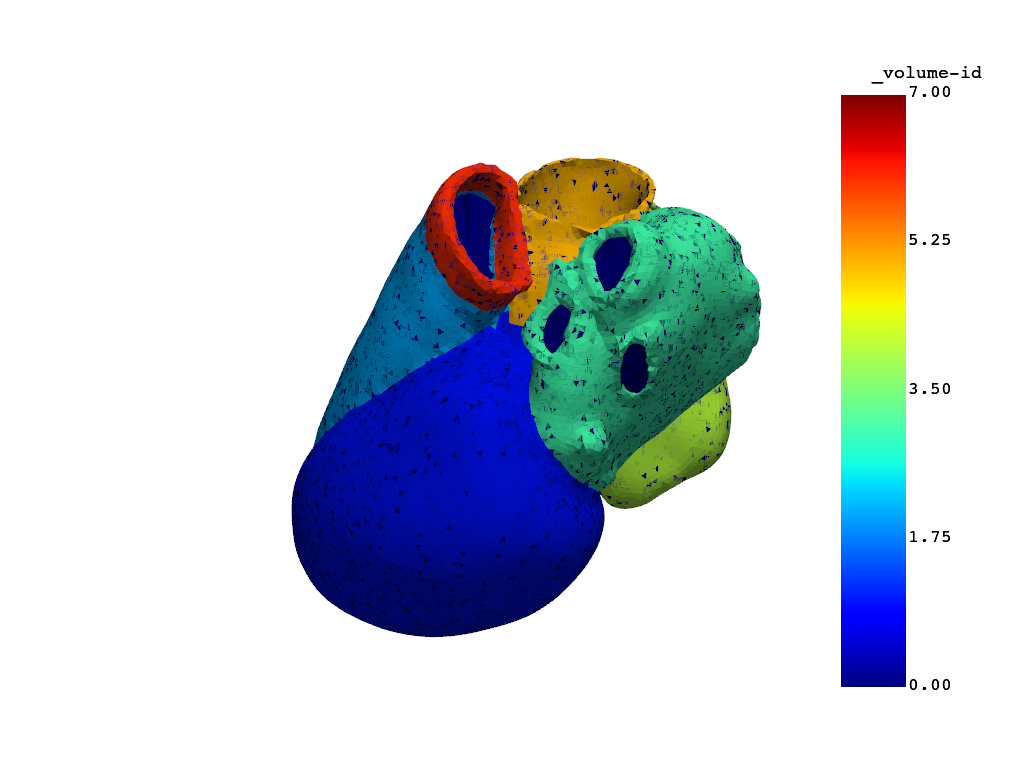
# Plot the remeshed model.
model.plot_mesh(show_edges=False)
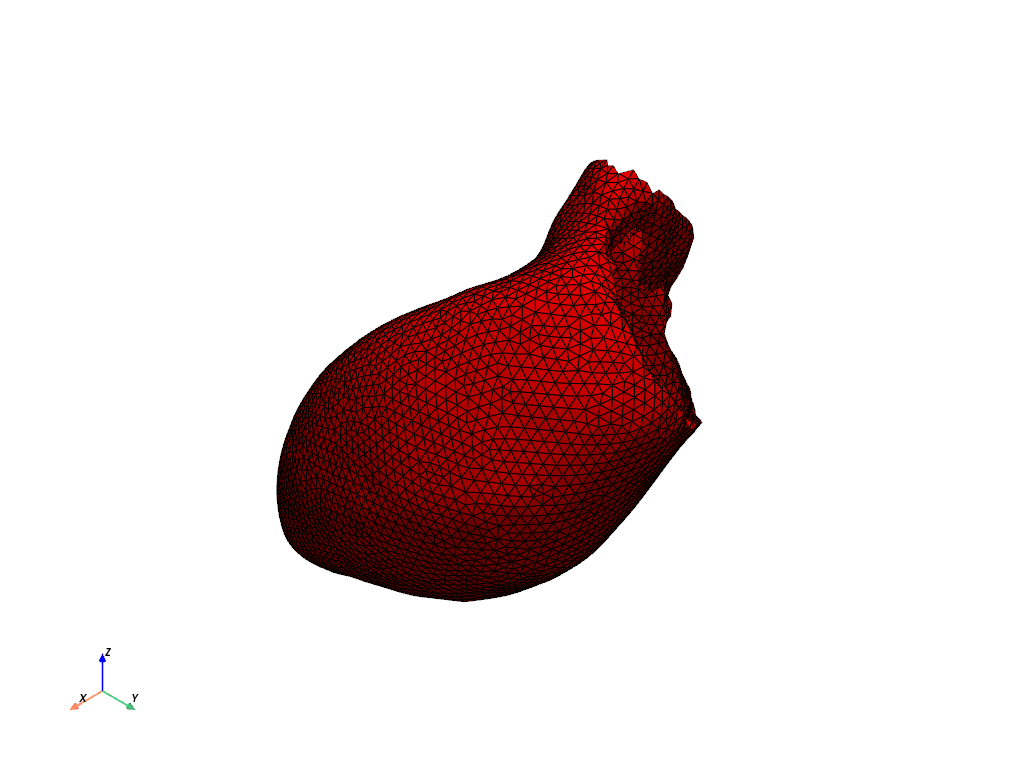
# Plot the endocardial surface of the left ventricle.
model.left_ventricle.endocardium.plot(show_edges=True, color="r")
# Use Pyvista to plot all cavity surfaces.
import pyvista as pv
all_cavities: pv.PolyData = pv.merge([c.surface for c in model.cavities])
all_cavities.plot(show_edges=True)


Volume of LV cavity: 120446.96049660796 mm^3
Volume of LV cavity: 68001.90309648326 mm^3
Total running time of the script: (7 minutes 36.912 seconds)

